
Stories
Aaron Gitler: Discoveries that Give Hope for ALS

Aaron Gitler began his scientific career as an undergraduate at Pennsylvania State University, where he volunteered in a lab washing dishes. Despite the less-than-glamorous work, he loved being there. Eventually, he realized that if he finished his dishwashing early enough, he could observe students and postdocs doing their research experiments. In time, they let him do experiments of his own. “In retrospect, none of the experiments really worked out,” says Gitler, “but I learned a lot and had a lot of fun. It prepared me for my next research experiences.”
After college, Gitler worked in Jonathan Epstein’s lab at the University of Pennsylvania. There he performed experiments and learned about embryonic heart development and congenital heart disease. He became so excited about this work that he applied to UPenn for graduate school and ultimately joined Epstein’s lab for his dissertation research. His project explored how the valves of the heart form and how blood vessels migrate. Gitler discovered several genes involved in the regulation of these processes and studied those genes in mice. “I loved it all,” Gitler remembers.
After receiving his doctorate from UPenn, Gitler started looking for a postdoc position. He applied to several developmental biology labs as well as to the lab of Susan Lindquist at the Whitehead Institute for Biomedical Research, in Cambridge, Massachusetts. Lindquist’s lab belonged to a different subfield—protein biology—but it always produced papers that Gitler enjoyed reading. “I wanted to go to that lab because I just thought the science was so creative and exciting,” says Gitler. “When I went to visit, I was completely blown away. Even though I didn’t know how to do any of the techniques they were working with, I thought that if I went there, good things would happen.” Gitler joined the Lindquist lab and was grateful he did. He has fond memories of Lindquist, who unexpectedly passed away in 2016. “She was an amazing mentor and a tremendously creative scientist,” he says.
As a postdoc, Gitler used a simple model system—yeast—to study protein misfolding and its role in Parkinson’s disease. In 2006, just as he was about to start his own lab at UPenn, a protein called TDP-43 was discovered that contributes to amyotrophic lateral sclerosis, or ALS. Gitler realized that he could apply the same approaches he had been using to study Parkinson’s to ALS. “It really worked out,” Gitler says, “and I’ve been pursuing it for 15 years.”
“ALS is a devasting disease, and it would be really meaningful if some of our work could translate into humans. That would just make my heart soar.”
Through his experiments on yeast, Gitler investigated how and why these misfolded proteins aggregate in the brains of people with ALS and how this process might be hindered. Using genetic screens, he overexpressed or deleted genes to find ones that might alter the course of the disease. One gene he identified through this process had a human homolog—a gene in yeast that is similar to the gene in humans—called ataxin-2. Normal ataxin-2 has 22 repeats of the nucleotide sequence CAG; when the number of repeats increases to more than 34, it triggers a degenerative disease of the nervous system called ataxia. Gitler was interested in exploring what happens when there is an intermediate number of repeats (i.e., between 27 and 33).
In 2009, as an assistant professor at UPenn School of Medicine, Gitler was selected as a Rita Allen Foundation Scholar, and he was able to delve into this question. Gitler collected DNA samples from more than 900 ALS patients for analysis and found that 5% of them had an intermediate number of repeats. “This was probably my biggest discovery, and it has really transformed my lab,” says Gitler. Not only was he able to show that a simple model system like yeast could be used for powerful applications and insights into human disease, but he also found a new gene associated with ALS.
Since this discovery, Gitler has made considerable progress in understanding ataxin-2. In 2017, his team—now at Stanford University—was able to replicate these experiments using a mouse model and showed that mice with the ALS gene TDP-43 lived longer when levels of the protein ataxin-2 were significantly reduced. Pharmaceutical companies have also shown interest in targeting ataxin-2 as an ALS therapeutic. Gitler is awed by this progress. “I hope it works,” he says. “ALS is a devasting disease, and it would be really meaningful if some of our work could translate into humans. That would just make my heart soar.”
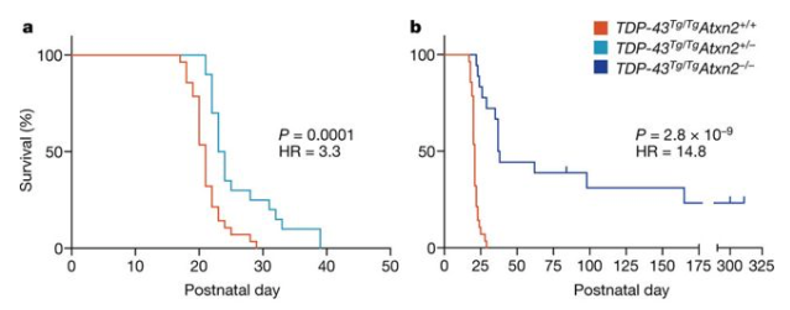
(Image: Becker, et al. [2017] Nature)
Gitler is now the Stanford Medicine Basic Science Professor of Genetics at Stanford University. Here, he reflects on his time as a Rita Allen Foundation Scholar, describes some of the research opportunities on the horizon, and offers advice to early-career researchers.
What was your experience like as a Rita Allen Foundation Scholar?
I’m extremely grateful for the support of the Rita Allen Foundation for two reasons. One, the funding at a very early stage allowed me to try risky strategies—studying a brain disease using yeast. Having this funding meant I didn’t have to listen to all the people telling me that this is too risky, that it isn’t going to work. I could just say, I have a few years of funding—I’m going to go for it. While some of my colleagues made more incremental advances because they had to play it safe, I was able to make larger ones. In that way, it’s really helped my career.
Two, I had access to the network of other Scholars, many of whom I still connect with as friends and colleagues. Their level of rigor and ambitiousness motivated me because I wanted to keep up with them, and that led me to work harder and try bolder approaches than I might have otherwise.
Looking toward the future, what new opportunities will the latest tools make possible?
When I moved my lab to Stanford in 2012, it was the beginning of CRISPR, and I thought, why don’t we see if we can use CRISPR to do the same screens that we’re doing in yeast, but in the human genome—all 20,000 human genes rather than just 6,000 yeast genes? We’ve made major progress in that, and the first papers are starting to come out. That’s really revolutionized the lab.
Another new area is single-cell sequencing. You can now sequence the RNA from a single cell, and then you can learn what type of cell it is and lots of other information. So that’s a powerful technology that we’ve been exploring. It’s important to stay up on the latest technologies, and I have to say, very few of my students are doing yeast experiments now, although we still have that in our corner.
The most exciting and also the most challenging part of running a lab is that you can’t rest on your laurels. You’re always having to take the next step.
What advice do you have for early-career scholars?
Three things: be open and giving; identify role models and try to emulate them; and separate people’s scientific successes and failures from their value as people.
When I became a senior grad student, I would look at the new postdocs and try to be like them. Then when I became a postdoc, I’d look to faculty members; and when I was an associate professor, I would look to the full professors.
I’ve learned a lot from others, including from my Ph.D. adviser, Jonathan Epstein. He would be very rigorous with everyone in the lab, but then outside the lab, when we would have social get-togethers, he would treat everyone completely equally, no matter how their project was going. This idea of separating a person’s value from whatever they have to do in the lab stuck with me, and I try to put it into practice in my own lab. From day to day, or year to year, some students’ projects, because of luck or other factors, may be doing better than others, but I don’t want any of them to feel that their value as a person depends on how some western blot turns out. I really am very grateful to all of my students. They work very hard. I care a lot about them, I am continuously inspired by each of them, and I try to help them in their careers.