
Stories
Hilary Coller: When Cells Sleep
Hilary Coller caught the biomedical research bug early—as a junior high school student she took part in research that resulted in a first-author scientific publication. Coauthored with her father Barry Coller, a blood and vascular biologist at The Rockefeller University, the paper explored a technical issue in making monoclonal antibodies, which are widely used as protein markers in biomedical research.

Hilary Coller
(Photo: Brian Wilson)
As her interests matured, Coller was drawn toward the policy dimensions of research. After completing her undergraduate degree in biochemistry and molecular biology at Harvard, she embarked on a master’s program in technology and policy at the Massachusetts Institute of Technology. She was interested in pursuing research that would be helpful to policymakers, so for her Ph.D. she joined the MIT laboratory of William Thilly, who was studying chemicals in the environment and their ability to cause cancer in humans. Her doctoral research explored the pattern of mutations such chemicals induced, focusing specifically on the DNA in the mitochondria, the power-supply organelles of the cell.
Having explored the external causes of mutations so intensively in her graduate work, Coller decided to spend her postdoc understanding cancer from a more cell-intrinsic perspective. She stayed on in Cambridge for a one-year fellowship at the Whitehead/MIT Center for Genome Research, where she worked with pioneering genomicists Eric Lander and Todd Golub (as well as cancer transcriptional regulation expert Robert Eisenman of the Fred Hutchinson Cancer Research Center in Seattle) to explore the functional pathway of the transcription factor and cancer driver Myc in regulating the cell cycle.
Next, she headed west for a second postdoc with Jim Roberts at the Fred Hutchinson Cancer Research Center, excited to apply the genome-wide analysis technologies she had learned to understanding the cell cycle and the causes of cancer. Cancer cells are characterized by rampant cell division and overgrowth—processes that are relatively well studied. Coller began to explore what happens when cells exit this proliferative cell cycle and become quiescent—not actively dividing, but maintaining their capacity to do so. This process is important in cancer, as its reversal may explain why tumors that have lain dormant, sometimes for years, reawaken to cause a recurrence of a patient’s cancer.
“He suggested that I consider applying microarray technology to a ‘dead-end’ cell cycle state of quiescence…I liked the idea of being a pioneer in studying a cell cycle state that was physiologically important but poorly understood.”
Coller’s early work on these sleeper cells suggested that, rather than passively lying inert, quiescent cells may be engaged in certain as-yet-unknown tasks. Quiescence, she and her colleagues proposed in a publication that laid out a novel view of the phenomenon, “was not simply a downstream consequence of exit from the cell cycle,” but an actively regulated process.
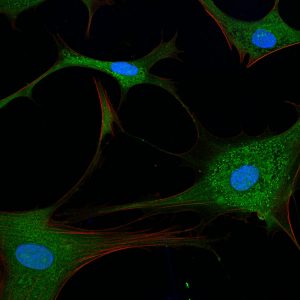
Rita Allen Foundation Scholar Hilary Coller uses fibroblasts, cells that form connective tissue in animals, to study cell quiescence, in which cells reversibly exit the cell cycle. Shown here are primary human fibroblasts visualized with fluorescent stains.
(Image: Sarah Pfau)
In 2005 Coller accepted an assistant professorship at Princeton University. She spent her very first day on the job in hot pursuit of high-quality paper to print her Rita Allen Foundation Scholars Award application. She raced to the FedEx drop-off box to meet the submission deadline, and was selected as an RAF Scholar later that year. She and her colleagues set out to understand the attributes of quiescent cells and the molecular mechanisms that regulate them. In 2010 they reported that quiescent cells can be as metabolically active as proliferating cells, despite the long-held assumption that they are napping on the job.
Interfering with the shift between quiescence and an active cell cycle could provide a therapeutic avenue for cancer treatment, Coller says. For example, it may be possible to prevent tumor resurgence by keeping the dormant cells in that state, or finding a way to target and kill them; another possibility is kicking them back into the cell cycle and targeting them with selective therapies.
Here, Coller recalls her early encounters with molecular biology and discusses the many remaining unknowns about cell quiescence.
What was your very first research experience?
In eighth grade, I remember learning about how a specific mutation in the gene encoding the sickle cell protein [a component of hemoglobin, which carries oxygen in the blood] results in sickle cell disease. I thought this was absolutely fascinating and got very excited about biomedical research. My first research experience was for a science fair project in ninth grade. My father was working with monoclonal antibodies and suggested I tackle a mathematical question about making monoclonal antibodies as a research project. I enjoyed working on the problem and presented it at the science fair. My father and I eventually published the work—it was my first scientific publication.
What drew you to studying quiescent cells, and what elements of the phenomenon has your lab focused on?
Early in my second postdoc, my advisor, Jim Roberts, pointed out to me that a lot of the important events in the cell cycle are post-translational modifications [at the protein level], and that these would be missed by microarrays, which only detect changes in gene expression. But he suggested that I consider applying microarray technology to a “dead-end” cell cycle state of quiescence. He said that not very much was known about this state, but that the cells have RNA, and would be amenable to transcriptomic analysis. I liked the idea of being a pioneer in studying a cell cycle state that was physiologically important but poorly understood.
One aspect of quiescence that we have been investigating is how quiescent cells reorganize their metabolic pathways. We’ve found that quiescent cells don’t necessarily get “sleepy” and shut down their metabolism, but can maintain high levels of metabolic activity. Further, while metabolic pathways are often considered unimportant or routine “housekeeping” functions of the cell, we’ve found that metabolic pathways can also have an important regulatory role. Understanding how and why quiescent cells reorganize their metabolic pathways has been a very interesting question for my laboratory.
What exciting directions is your lab currently pursuing?
A major focus for my laboratory for the past two years has been developing tools and methods for understanding quiescent versus proliferative cells in vivo using mouse models. This has been driven by the increasing availability of tools to visualize or monitor cell biological processes and manipulate specific pathways in intact organisms. We are also very focused on making connections between disease states and our basic research observations about quiescence in the laboratory. For example, we are establishing the infrastructure to ask whether the pathways that we found to be important for the transition between cellular proliferation and quiescence are relevant for patients who fail to mount a proper wound healing response, which is an important public health problem.