
Stories
Christopher Lima: Structure Meets Function
As an undergraduate at The Ohio State University, Christopher Lima split his time between studying biochemistry and playing the trombone in a ska band.
When the time came to choose between these two pursuits, Lima opted for a career in science. He delved into structural biology, first as a graduate student with Alfonso Mondragón at Northwestern University, where he mapped the doughnut-like structure of a bacterial topoisomerase enzyme to shed light on how it breaks and reseals DNA.

Christopher Lima
(Photo: Memorial Sloan Kettering Cancer Center)
Lima then conducted postdoctoral research with Wayne Hendrickson at Columbia University, where he crystallized two divergent members of the cryptic histidine triad (HIT) protein family. Emerging genomic sequences showed that the proteins were highly conserved among organisms from bacteria to humans, but their functions were not well understood.
The proteins’ structures suggested that they might transfer or remove nucleotides from other molecules. Years later, this work “came back to haunt me in a very satisfying way,” Lima says, as his colleagues found that a HIT protein called DcpS was responsible for removing the molecular “caps” of partially degraded messenger RNA molecules, thus preventing their interaction with the translation apparatus. Lima and colleagues then went on to resolve the structure of a DcpS-cap complex.
Mechanisms of RNA processing have since become a major area of focus for Lima, who established his own research group at Weill Cornell Medical College in 1998, and is now an Investigator of the Howard Hughes Medical Institute at Memorial Sloan Kettering Cancer Center. Lima and his team examine fundamental cellular processes that determine the fates of proteins and RNAs—with implications for the regulation of the cell cycle, gene expression and other vital activities.
Lima became a Rita Allen Foundation Scholar in 2003. At the time, small ubiquitin-like modifier (SUMO) proteins had recently been discovered. Researchers knew that these little tags could be attached to larger proteins, and that their presence was required for cells to properly grow and divide. But the specific targets of SUMOs, and even the mechanisms of their attachment, were unclear.
“This enabled me to do things that I couldn’t necessarily do with more traditional funding sources. To get that boost when you’re probably the least sure of what you should be doing to have a successful career…having that level of guidance was really a turning point.”
Structural studies by Lima and his group helped to illuminate the interactions between SUMOs and the enzymes that link them to other proteins—interactions critical for cell growth, stress responses and other functions.
Lima also had a longstanding interest in the molecular processes that generate and modify RNAs, which form diverse structures that mediate many aspects of the transformation from genetic information into actual cellular activities. This interest led Lima to explore the stunning complexity of the dozen or more proteins that make up the RNA exosome, which has the ability to either chew up RNA molecules or to convert them into mature, functional forms. How the exosome discriminates between these possibilities remains unclear.
Here, Lima reflects on the enigmas of the exosome, the future of structural biology and the intrinsic artistry of science.
Why was the exosome so difficult to decipher? How did you do it?
When we started working on the exosome, it wasn’t even clear which proteins were in it, or how many proteins or what the stoichiometry was. It turned out that the complex is dynamic: it has nuclear forms, it has cytoplasmic forms, and it has all sorts of different cofactors and accessory factors.
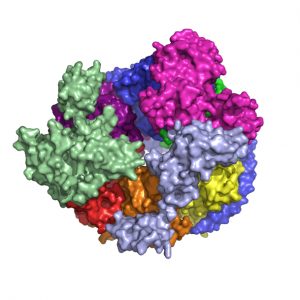
A model representing the architecture of the human RNA exosome core, which consists of nine distinct protein subunits, as elucidated by 2003 Rita Allen Foundation Scholar Christopher Lima and his team
(Image: Christopher Lima)
We realized that it was almost going to be an intractable problem until we were able to reconstitute it in vitro from individual components—which we did, more than a decade ago now. It was a landmark example of a brute force effort, in the sense that we tried basically every combination of every possible factor in order to get these complexes to form. But we succeeded.
We characterized the activities of our reconstituted samples, and showed that in many cases they were not as anticipated. We were then able to genetically modify individual components, rather than modifying the organism, to generate an intact complex with modified components. That was very critical, because most of the subunits are essential for viability. By looking in vitro, we were able to see what the individual subunits were actually doing.
Where are you going next with the exosome?
Now the big question is how it’s regulated. In some ways, it’s analogous to the proteasome, which everybody looked at as a trash can: you put ubiquitin on the protein, it finds the proteasome, and the proteasome just degrades—it has no other option. But it turns out there are multiple levels of regulation even at the site of degradation, and I suspect very strongly that that is also going to be true for the RNA decay system.
Several projects in my lab right now are focused on the question of how fate is decided once RNA reaches the exosome. When you make a particular structured RNA or structured piece of nucleic acid and you screw that up, how do you sense that it’s screwed up? How is that molecule labeled as screwed up, and then how does it get to the exosome to be interrogated?
I think that that level of quality control is something we understand at the genetic level right now. But at the mechanistic level, we have no clue.
What’s next for the field of structural biology?
The methodology defined the field for quite some time, and it still does in some cases. But now it’s the biology that’s driving the bus.
The big challenge is to become biologists who use structure, rather than structural biologists who explain other scientists’ biology with structure. As our systems are growing in complexity, this is becoming the major source of effort—not so much figuring out which structure to solve, but figuring out how the system behaves as a whole.
There are tens of factors that have to interact with one another to decide the fate of a particular RNA or protein. And that’s way more complicated than it was when I started in science. So it’s really about understanding the biology deeply enough.
Looking back at your career choice between science and the arts, how do you feel about that decision now?
At the time, I wasn’t quite aware of how creative science can be, if you’re lucky enough to have the opportunity. When I realized that the level of creativity I was employing in these other parts of my life could be applied to my science, that’s where it really took off. Now I spend most of my time trying to think of new ways to get at the problems we’re investigating, and that still flexes those creative muscles.
I don’t view science as a job—it’s more of a lifestyle. If I couldn’t come into my office, I’d be sitting at home thinking about the same stuff. It’s really a privilege.